We are interested in a diversity of topics at the intersection of ecology and evolution, with a special focus on fungal biology. Our work in mycology reaches across boundaries with respect to microbiology, plant pathology, genomics, functional traits analysis, and phylogenetics.
Here are some of our active projects and ongoing interests in plant microbiomes, plant health, fungal biodiversity, and molecular ecology. For more about our work through the Robert L. GIlbertson Mycological Herbarium, please click here. Para obtener información en español sobre las huellas moleculares de animales silvestres, haz clic aqui.
Phylogenetic diversity and the origins of plant-fungal symbioses
in the most species-rich fungal phylum
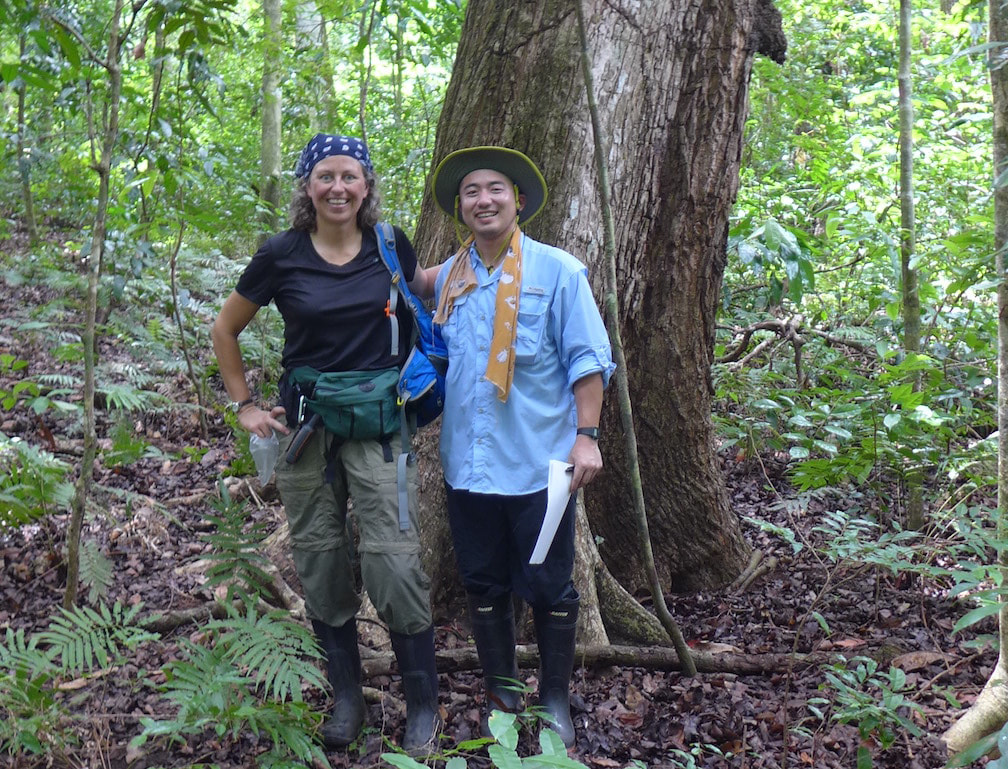
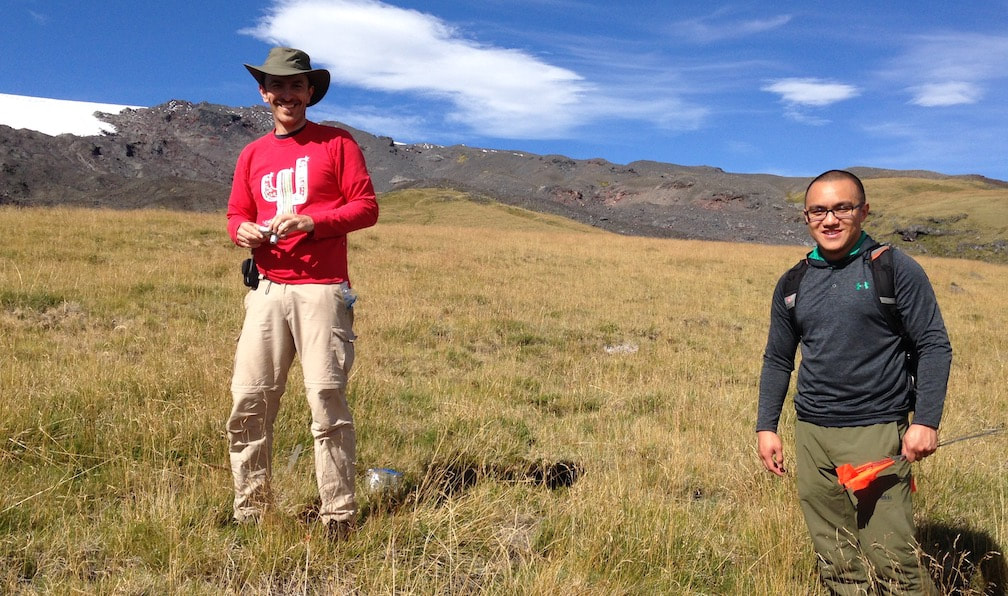
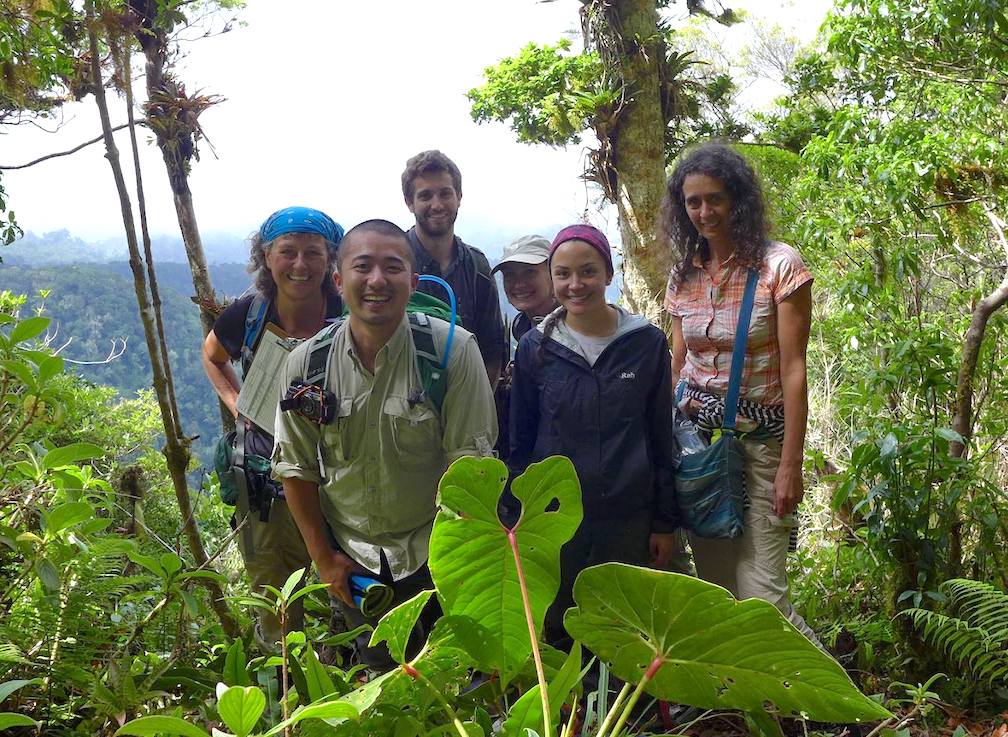
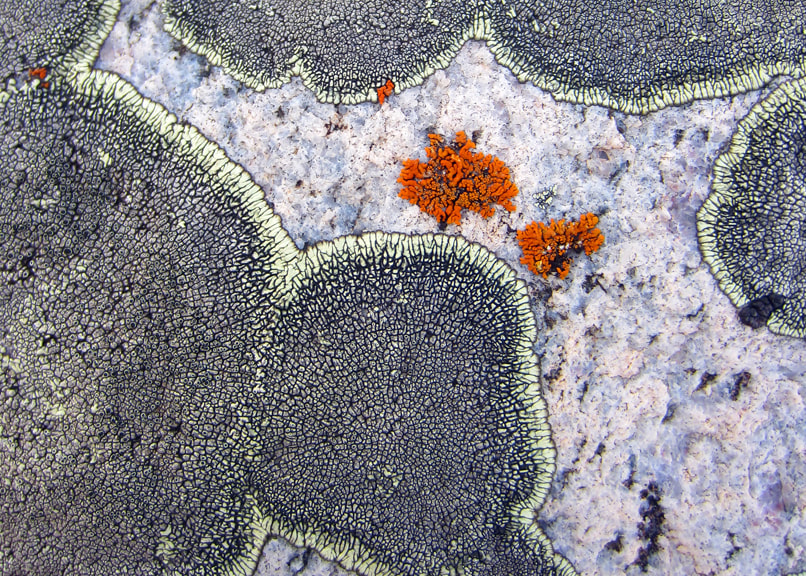
Why are plant-fungal associations ubiquitous in all terrestrial ecosystems? How did the fungal component of leaf microbiomes arise and diversify? How can endophytic fungi fill gaps in the fungal tree of life, thus informing our perspective on evolutionary dynamics in the most diverse fungal lineages? These questions frame our collaboration with lead researchers at Duke University (François Lutzoni and Jolanta Miadlikowska) and collaborators at UConn (Louise Lewis), Ole Miss (Erik Hom), NC State University (Ignazio Carbone), and overseas universities in Chile, Poland, Hungary, and Belgium. Fieldwork in diverse ecosystems located in global biodiversity hotspots is the centerpiece of a multi-level study that ranges from the associations of fungi with cyanobacteria and algae to phylogenomics of the major lineages of fungi. We are grateful to be collaborating with researchers in each of our major field sites, including areas of Panama, Chile, Borneo, and South Africa, and with global experts in lichenology, algal biology, soil fungi, and bioinformatics. Funded by the NSF Genealogy of Life program (GoLife).
Here are some of our active projects and ongoing interests in plant microbiomes, plant health, fungal biodiversity, and molecular ecology. For more about our work through the Robert L. GIlbertson Mycological Herbarium, please click here. Para obtener información en español sobre las huellas moleculares de animales silvestres, haz clic aqui.
Phylogenetic diversity and the origins of plant-fungal symbioses
in the most species-rich fungal phylum
Why are plant-fungal associations ubiquitous in all terrestrial ecosystems? How did the fungal component of leaf microbiomes arise and diversify? How can endophytic fungi fill gaps in the fungal tree of life, thus informing our perspective on evolutionary dynamics in the most diverse fungal lineages? These questions frame our collaboration with lead researchers at Duke University (François Lutzoni and Jolanta Miadlikowska) and collaborators at UConn (Louise Lewis), Ole Miss (Erik Hom), NC State University (Ignazio Carbone), and overseas universities in Chile, Poland, Hungary, and Belgium. Fieldwork in diverse ecosystems located in global biodiversity hotspots is the centerpiece of a multi-level study that ranges from the associations of fungi with cyanobacteria and algae to phylogenomics of the major lineages of fungi. We are grateful to be collaborating with researchers in each of our major field sites, including areas of Panama, Chile, Borneo, and South Africa, and with global experts in lichenology, algal biology, soil fungi, and bioinformatics. Funded by the NSF Genealogy of Life program (GoLife).
How do functional traits of leaves predict and respond
to functional traits of foliar endophytes?
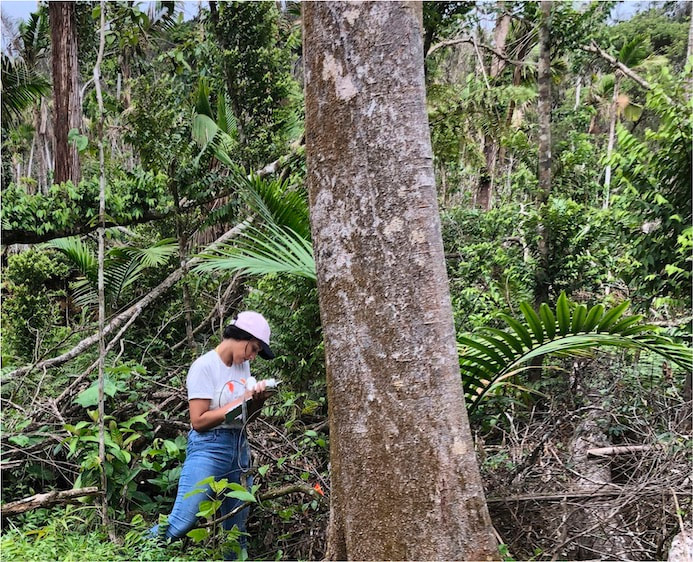
The worldwide Leaf Economic Spectrum (LES) describes functional traits of leaves of plants worldwide, integrating life history, evolutionary tradeoffs, structural traits, and leaf defenses. How do foliar endophytes reflect and extend those functional traits? With colleagues at Tulane University (project leader, Sunshine Van Bael, team members in her lab, Peter Tellez, Mareli Sanchez Julia, Bolivar Aponte Rolón), we are collaborating to study endophytes in the world's most species-rich tropical ecosystems -- lowland tropical forests. Our work adds an endophyte dimension to the LES and links functional traits of both leaves and fungi across diverse tropical species. Funded by NSF, DEB - Population & Community Ecology
Seed-associated fungi in tropical forests:
keys to forest dynamics
The tremendous diversity of tree communities in tropical forests is maintained in part through density-dependent mortality imposed by pathogens and herbivores. Most of our understanding of these processes focuses on seedlings and saplings, but the strategies by which seeds are defended, and the roles that seed-associated fungi might play in shaping forest dynamics, remain less well understood. In a project led by Jim Dalling (University of Illinois), Camilo Zalamea (STRI/University of South Florida), Adam Davis (UI), and Carolina Sarmiento (USF), we are examining the factors that structure communities of seed-associated fungi and how such fungi affiliate with, and differentially impact, seeds of various species and dormancy classes. Funded by NSF, DEB - Population & Community Ecology
Tracking temporal and spatial shifts of endophytic symbionts
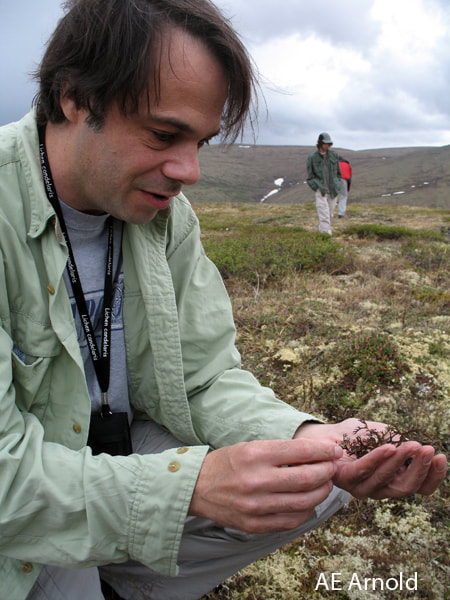
in the threatened terrestrial Arctic
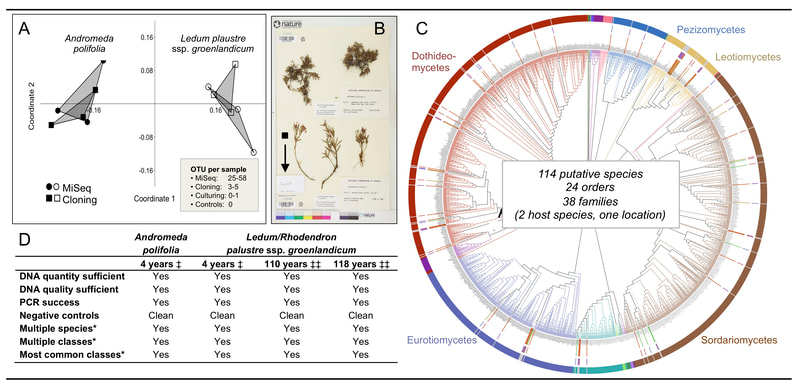
The terrestrial Arctic is imperiled by rapid climate change. How will fungal symbionts of iconic plants and lichens in the Arctic respond as the climate shifts? This UA-based project (with co-PI Jana U'Ren) is a collaboration with Barnabas Daru (TAMUCC), François Lutzoni and Jola Miadlikowska (Duke University), and Ignazio Carbone (NCSU) to study temporal, geographic, and functional shifts in endophyte communities relevant to plants and lichens in the North American Arctic. Our temporal framework comes from our recent discovery that herbarium specimens of plants and lichens can serve as 'time capsules' of symbiont communities at the time of collection. Revisiting the sites as many as 200 years after those collections gives us an opportunity to track changes in host- and symbiont distributions. Funded by NSF, DEB - Systematics and Biodiversity Science
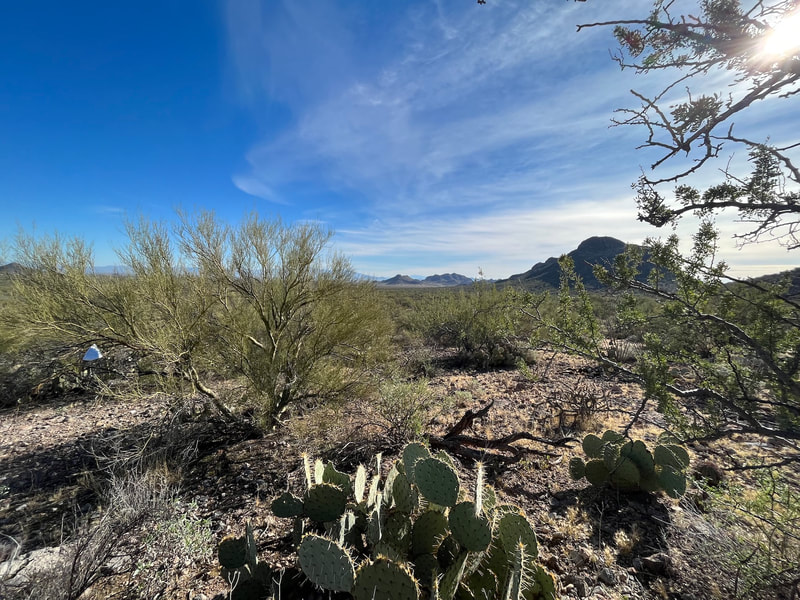
Plant-fungal symbioses in a changing world:
perspectives from arid, semi-arid, and tropical landscapes
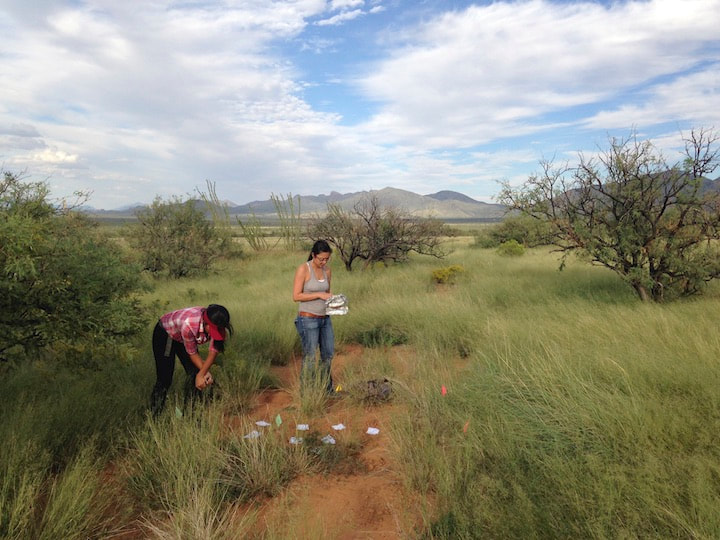
We are interested in how a changing environment effects the fungal communities that are integrally linked with their hosts. Our projects in this area focus on how fungal endophytes and ectomycorrhizal fungi, and their associations with plants, shift under altered regimes of fire (in the Sky Islands and grasslands of Arizona), land use (southern Arizona), plant invasions (as by Lehmann's lovegrass in southeastern Arizona, and buffelgrass in the Sonoran Desert), and climate disturbances (such as hurricanes in Puerto Rico, drought in Arizona, and continental-scale climate dynamics observed through a network of NEON and LTER sites)... and more. These diverse projects are student-led and funded by diverse sources. Contact us to learn more! Funded by NSF, Macrosystems Research Award
Understanding genomic, evolutionary,
and symbiotic drivers of fungal phenotypes
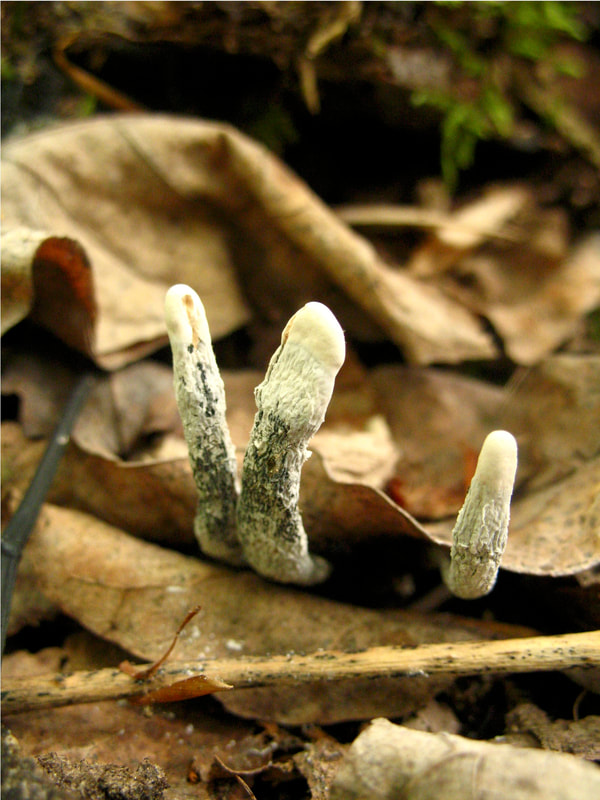
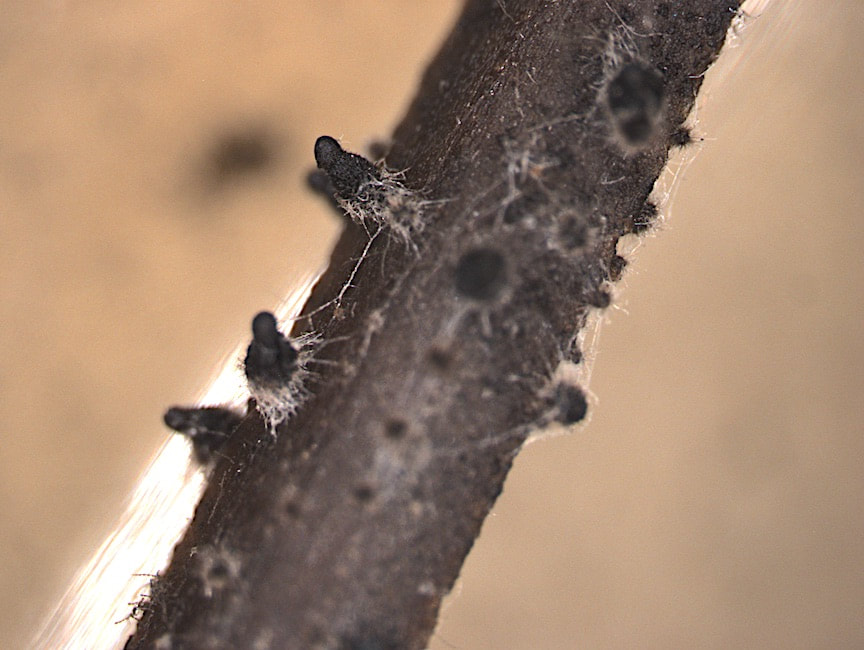
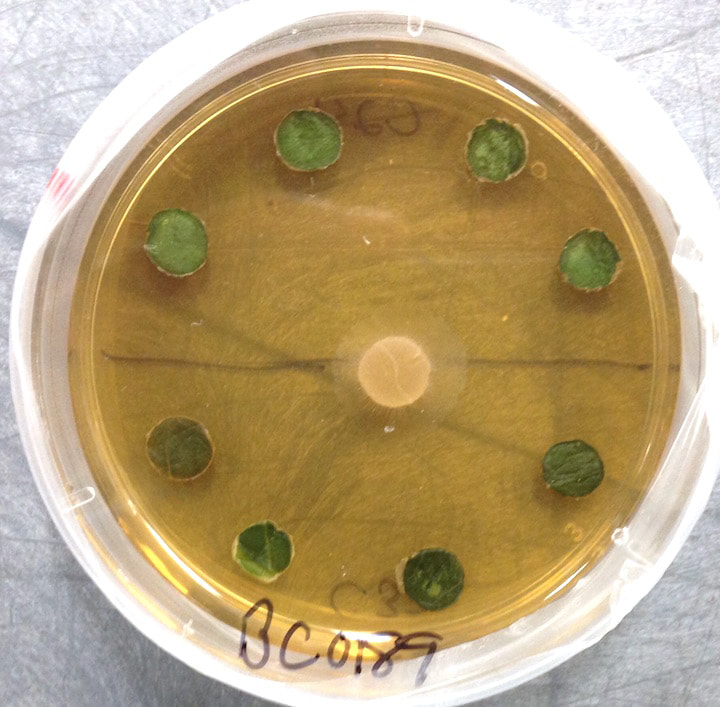
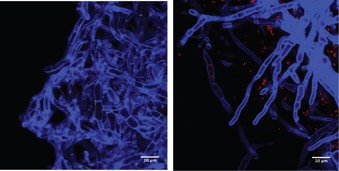
How labile are avirulent associations between plants and their hosts? To what degree do closely related fungi differ in their ecological modes? And what factors drive trophic mode transitions in fungi over ecological and evolutionary time? We ask these questions through diverse lenses. Our lab leads and co-leads projects to understand how endohyphal bacteria of fungal endophytes influence substrate use and ecological functions in vitro and in the field (collaborator: Dave Baltrus, UA). In projects led by Jana U'Ren (UA) we study fungal genomes, especially in Xylaria and related fungi, with collaborative projects with industry partners helping us discover the genomic underpinnings of symbioses. Finally we use inoculation experiments to understand how extrinsic conditions influence the outcome of plant-fungal interactions. These projects are funded by diverse sources (NSF IOS, DOE-JGI, and private sources).
Understanding genomic, evolutionary,
and symbiotic drivers of fungal phenotypes
How labile are avirulent associations between plants and their hosts? To what degree do closely related fungi differ in their ecological modes? And what factors drive trophic mode transitions in fungi over ecological and evolutionary time? We ask these questions through diverse lenses. Our lab leads and co-leads projects to understand how endohyphal bacteria of fungal endophytes influence substrate use and ecological functions in vitro and in the field (collaborator: Dave Baltrus, UA). In projects led by Jana U'Ren (UA) we study fungal genomes, especially in Xylaria and related fungi, with collaborative projects with industry partners helping us discover the genomic underpinnings of symbioses. Finally we use inoculation experiments to understand how extrinsic conditions influence the outcome of plant-fungal interactions. These projects are funded by diverse sources (NSF IOS, DOE-JGI, and private sources).
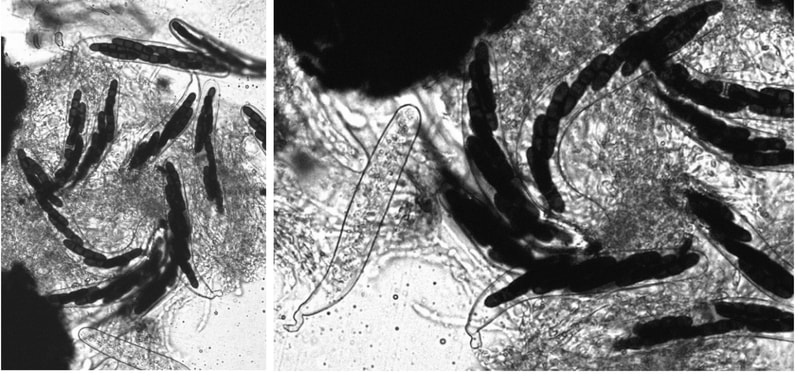
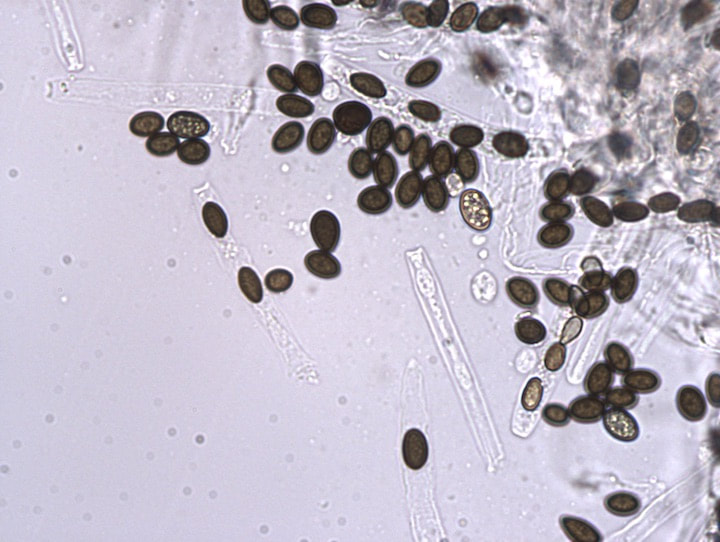
Discovering and describing fungal diversity
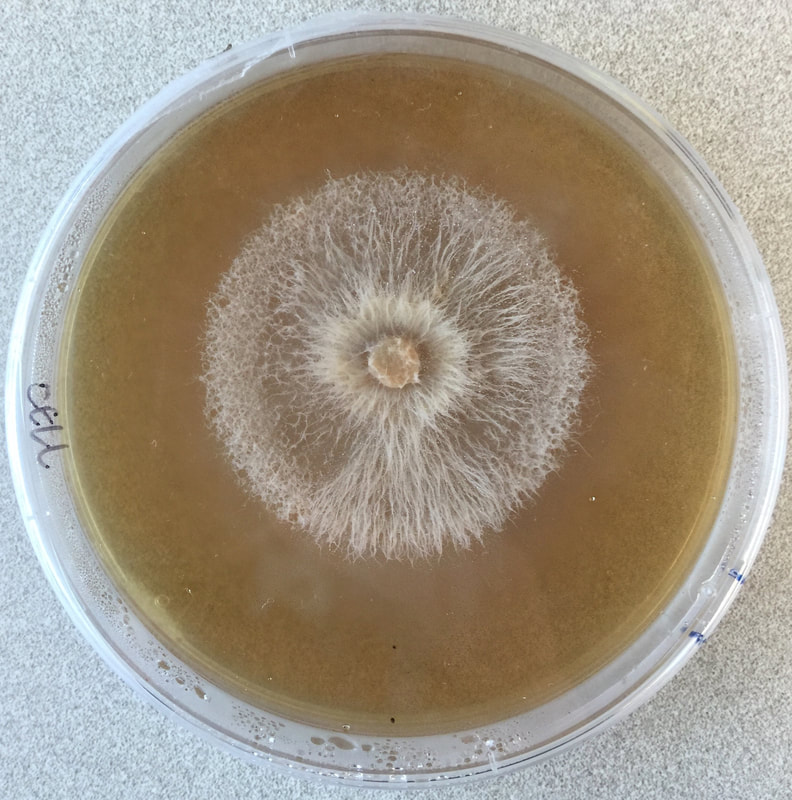
One of the great challenges in fungal biology is to link ecological studies, typically based only on barcode sequences, to the grand history of fungal systematics and taxonomy. We are exploring this through the lens of endophytes, describing novel species and thus expanding not only the fungal tree of life, but also our understanding of the ecology and evolution of already known groups. We have a special fondness for Coniochaeta, one of the most common genera in our endophyte collections in temperate and boreal forests, and Preussia, especially common in desert gnetophytes and angiosperms. These projects are funded by diverse sources (NSF IOS, DOE-JGI, and private sources).
Discovering and describing fungal diversity
One of the great challenges in fungal biology is to link ecological studies, typically based only on barcode sequences, to the grand history of fungal systematics and taxonomy. We are exploring this through the lens of endophytes, describing novel species and thus expanding not only the fungal tree of life, but also our understanding of the ecology and evolution of already known groups. We have a special fondness for Coniochaeta, one of the most common genera in our endophyte collections in temperate and boreal forests, and Preussia, especially common in desert gnetophytes and angiosperms. These projects are funded by diverse sources (NSF IOS, DOE-JGI, and private sources).
:
Molecular ecology of animals, their DNA 'fingerprints',
and their microbiomes
Many of the tools we use can be translated from plants for inquiries relevant to wildlife ecology. We are especially interested in passive sampling methods that allow us to learn about animals without perturbing them directly. We are studying how guano from endangered bats can help us estimate population size and movements across the landscape of the border region (a collaborative project with Bob Steidl, UA SNRE); how shed feathers can identify nests of cryptic songbirds in grasslands or population structure in owls; how feces can help us determine the presence of endangered animals in protected lands; how the oral microbiomes of urban hawks in Tucson changes as individuals mature; and how lizards' cloacal microbiomes may provide protection for eggs (a study led by Stacey Weiss, University of Puget Sound). Contact us for more info on these diverse projects. En español
Molecular ecology of animals, their DNA 'fingerprints',
and their microbiomes
Many of the tools we use can be translated from plants for inquiries relevant to wildlife ecology. We are especially interested in passive sampling methods that allow us to learn about animals without perturbing them directly. We are studying how guano from endangered bats can help us estimate population size and movements across the landscape of the border region (a collaborative project with Bob Steidl, UA SNRE); how shed feathers can identify nests of cryptic songbirds in grasslands or population structure in owls; how feces can help us determine the presence of endangered animals in protected lands; how the oral microbiomes of urban hawks in Tucson changes as individuals mature; and how lizards' cloacal microbiomes may provide protection for eggs (a study led by Stacey Weiss, University of Puget Sound). Contact us for more info on these diverse projects. En español
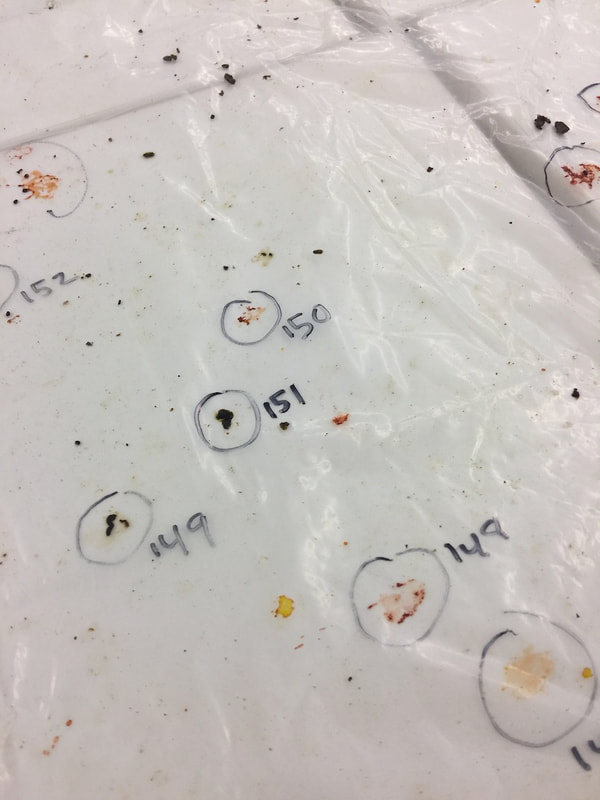
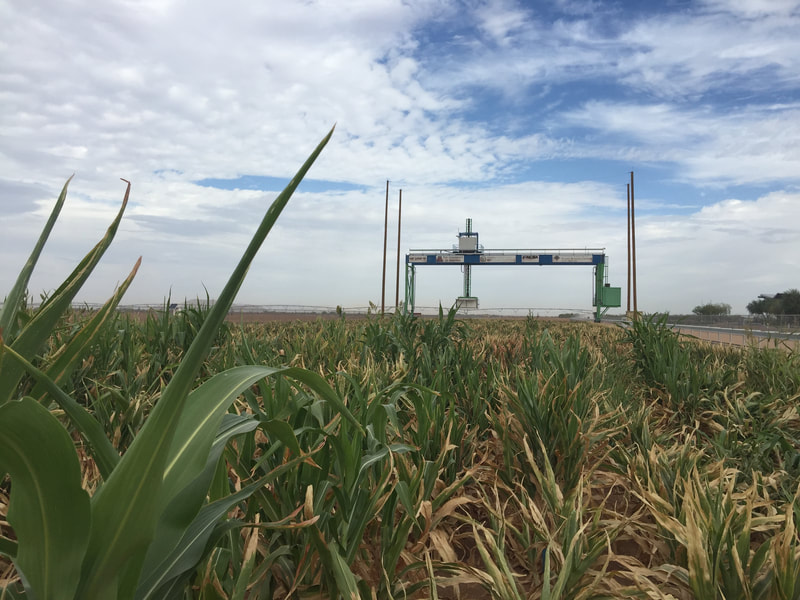
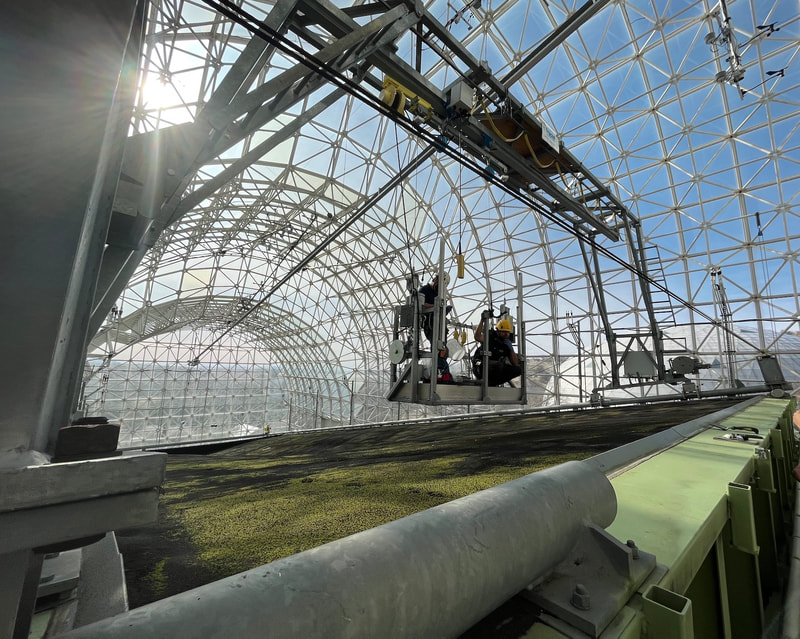
Plant microbiomes in arid lands: wild relatives of crop plants
and Arizona's remarkable biodiversity
We study diverse microbes associated with desert plants to understand how they provide or promote key traits for plant productivity and plant health. Our studies range from cultivated sorghum and lettuce at the Maricopa Agricultural Center (in collaboration with Duke Pauli) to soil-borne and seed-associated fungi as tools for ecological restoration in damaged grasslands, to the translation of microbiomes from wild relatives of lettuce and cotton to cultivated crops, to the diversity of endophytic fungi in unusual and dominant desert plants, to the establishment of biocrusts and their maintenance in wild systems and at the Landscape Evolution Observatory of Biosphere2. We connect these efforts with land managers and community scientists. Click here to learn more.